 Today you will use SDS-PAGE to visualize the effectiveness of IPTG induction and the purification procedure. In addition, the flow-through from the wash steps was stored. Part 2: Visualize purified protein with polyacrylamide gel electrophoresis (PAGE)ĭuring the previous laboratory session, you reserved an aliquot of your IPTG-induced and the uninduced cell lysate. Prerna Bhargava, will join us today for a workshop on writing impactful abstracts and titles. 
Protocols Part 1: BE Communication Lab workshop Because absorbance and protein concentration have a linear relationship, it is possible to compare the absorbance of an unknown protein sample to a standard curve, generated with samples of known protein concentrations, and calculate the concentration of protein in the experimental sample.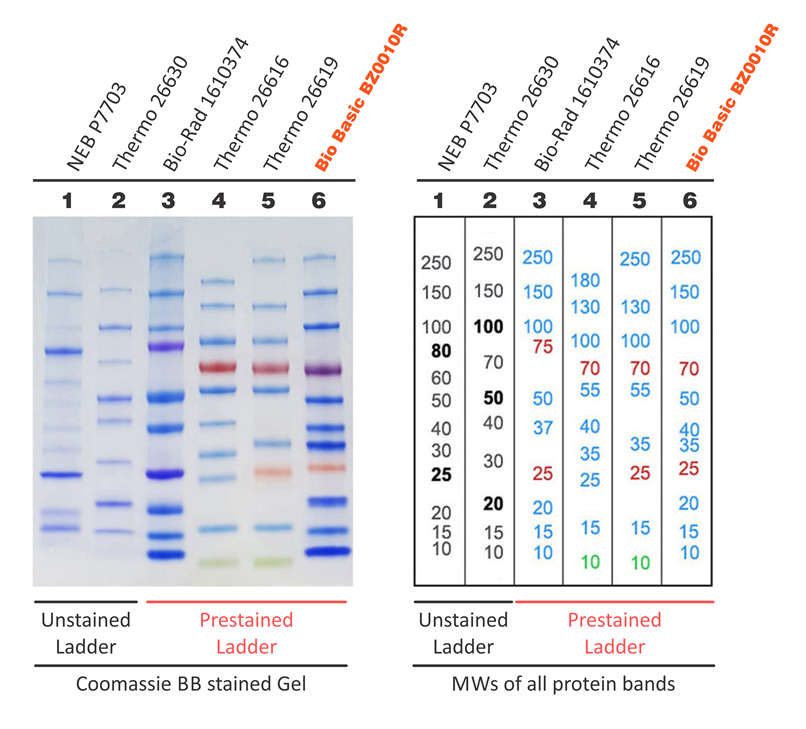 
This water-soluble complex has a strong absorbance at 562 nm. Through this reduction reaction, a purple product is formed by the chelation of BCA and Cu 1+ at a 2:1 ratio. The Cu 1+ is formed when Cu 2+ is reduced by protein in an alkaline environment. The ability to measure protein concentration is based on the detection of Cu 1+ by the detection reagent, bicinchoninic acid (BCA). This kit enables colorimetric detection and quantification of the total protein within a sample. To measure the concentration of your purified protein, you will use the BCA Protein Assay Reagent Kit. Because this is a non-specific stain for all proteins, it will provide information concerning the purity of your protein sample. To visualize all the proteins released by the bacteria, you will stain the gels with Coomassie Brilliant Blue (actually, a variant called BioSafe Coomassie). When running –IPTG and +IPTG samples side-by-side, you should see a protein band at the expected molecular weight for FKBP12 (~14 kDa + 3 kDa from the His-tag = 17 kDa), which may be very faint or non-existent in the -IPTG control sample, but bright and thick in the +IPTG induced sample. (Non-denatured proteins run according to their molecular weight, shape, and charge.) You will run a reference ladder containing proteins of known molecular weight and amount to determine the size and concentration of your purified FKBP12 protein. Since denatured proteins are linear, they will move through the gel at a speed inversely proportional to their molecular weight, just like DNA on agarose gels. SDS is an ionic surfactant (or detergent), which denatures the proteins and coats them with a negative charge. You will use sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) to evaluate your purified protein. Over time fragments of similar length accumulate into “bands” in the gel. 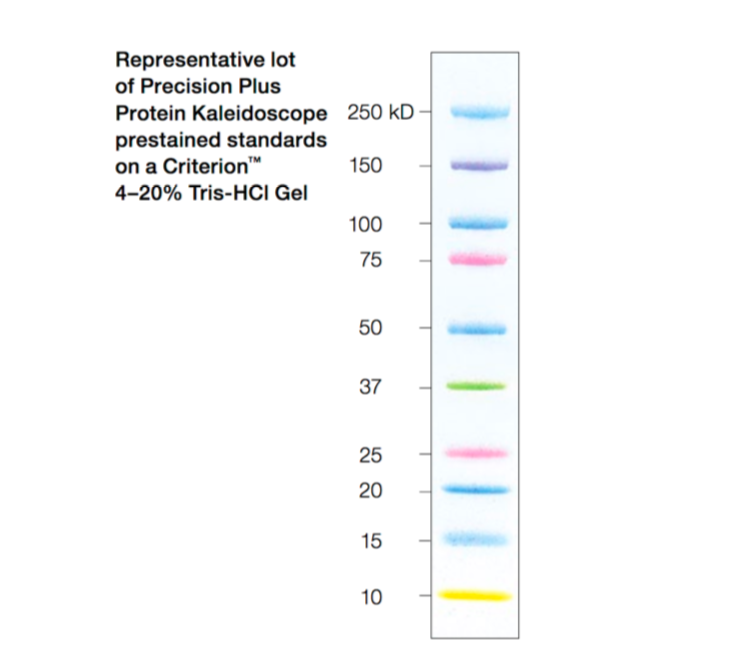
The distance a nucleic acid or amino acid fragment travels is inversely proportional to the log of its length. The larger molecules get entwined in the matrix and are stalled the smaller molecules wind through the matrix more easily and travel farther away from the well. The molecules to be separated enter the matrix through a well at one end and are pulled through the matrix when a current is applied across it. DNA, RNA and proteins are the molecules most often studied with this technique agarose and acrylamide gels are the two most common sieves. 2.4 Part 4: Concentrate FKBP12 protein solutionĮlectrophoresis is a technique that separates large molecules by size using an applied electrical field and a sieving matrix.2.3.2 Part 3b: Prepare Working Reagent (WR) and measuring protein concentration.2.3.1 Part 3a: Prepare diluted albumin (BSA) standards.2.3 Part 3: Measure protein concentration.2.2 Part 2: Visualize purified protein with polyacrylamide gel electrophoresis (PAGE).2.1 Part 1: BE Communication Lab workshop. 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
